While at Insight, I had the opportunity to create a data science project that demonstrates my ability to perform an analysis that creates business value within a short timeframe (about three weeks), simulating how data science operates in an industry setting. For this project I consulted with Zendo Tools, a company with an app that helps to track a user’s meditation progress.

The Zendo Tools app is used in conjunction with an Apple watch to record the user’s meditation session. Afterwards, the app shows how the person’s heart rate, heart rate variability, and motion changed throughout the session. It also compares these metrics across all of the meditation sessions over time. Heart rate variability is a metric of particular interest because it is known to be associated with a variety of measures of health and well-being and can potentially be improved with meditation.
Zendo Tools was interested in creating a product feature that reports to users when they were in the meditative state during their meditation sessions. This additional feedback would be useful because it gives users more information that can be used to improve on subsequent meditation sessions. I was provided with a couple hundred meditation sessions from a single user in order to perform this analysis. There were no labels of when the meditative state was reached in these sessions.
Before I began, I knew there would be several challenges:
- The meditative state does not have a concrete definition. How will I go about classifying this?
- Data is only coming from one user. I need to make sure the classification is generalizable to any user.
- The data only has three features (all time-varying). Will this be enough information to discriminate the meditative state?
My plan was to start with an unsupervised learning approach and see if I am able to obtain any meaningful clustering. Due to the limitations of the data noted above, it is reasonable to expect that this method may not succeed. If that is the case, I will instead manually create a decision rule for determining the meditative state.
Some key steps of the data preprocessing involved feature extraction and the creation of subsequences within each of the time series data sets. Using the three starting features, new features were created based on (1) the difference from baseline, (2) a rolling mean (e.g. the mean heart rate at a point in time +/- 15 seconds), and (3) a rolling percent change (e.g. the percent change in heart rate relative to 30 seconds ago). The following Python code demonstrates the process of creating the subsequences, in this case continually shifting over 1 measurement and storing a 3-measurement-long window.
segment_len = 3
slide_len = 1
segments = []
for i in df_list:
for start_pos in range(0, len(i), slide_len):
end_pos = start_pos + segment_len
segment = np.copy(i[start_pos:end_pos])
if len(segment) != segment_len:
continue
segments.append(segment)
After the data was preprocessed and divided into subsequences, I attempted two different clustering techniques: K-means Clustering and Gaussian Mixture Models. K-means clustering seemed like a logical starting point since it is simple and commonly used in this time series application. I also assessed the Gaussian Mixture Models algorithm since it overcomes a key limitation of K-means clustering, which is that clusters must be circular/spherical in shape.
With either method, I found that the clusters were not distinct based on the elbow method and Silhouette analysis. It also appeared that the clusters were not particularly meaningful based on inspection of their descriptive statistics. The code and plot below shows that the K-means elbow plot never reaches a clear elbow, indicating that there is no obvious number of distinct clusters.
from sklearn.cluster import KMeans
distortions = []
for i in range(1, 9):
km = KMeans(n_clusters=i, random_state=0)
km.fit(segments_array_2d)
distortions.append(km.inertia_)
plt.plot(range(1, 9), distortions, marker='o')
plt.title('K-means Elbow Plot')
plt.xlabel('Number of clusters')
plt.ylabel('Distortion');
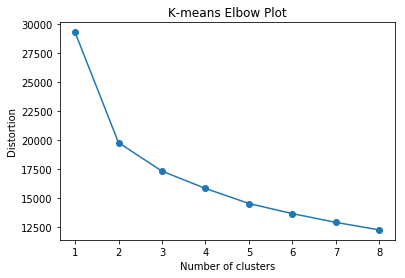
So ultimately, the best approach was to create a manual decision rule. I made these rules with guidance from Zendo Tools and based on papers found in the journal Applied Psychophysiology and Biofeedback. The criteria is as follows:
During the time window, the user must:
- Not be in motion
- Have no recent significant motion changes
- Have no recent significant heart rate changes
- Not have a significantly elevated heart rate
- Have a heart rate variability above baseline
As the code below demonstrates, these criteria are more strict during the early stages of the meditation session. Also, the meditative state is impossible during the very beginning of the session.
df_meditation_list = []
for index, i in enumerate(df_list):
if index < 13:
continue
if index > 120:
if ((i.motion > .2).any() or (i.motion_change > .2).any() or (i.sdnn_diff_bl < 0).any()
or (i.hr_diff_bl > .95).any() or (i.hr_pct_change > .95).any() or (i.hr_pct_change < .05).any()):
continue
else:
df_meditation_list.append(i)
else:
if ((i.motion > .2).any() or (i.motion_change > .2).any() or (i.sdnn_diff_bl < 10).any()
or (i.hr_diff_bl > .75).any() or (i.hr_pct_change > .75).any() or (i.hr_pct_change < .25).any()):
continue
else:
df_meditation_list.append(i)
The graph below shows how this criteria would classify an example meditation session.

To wrap it all up, I provided Zendo Tools with my Python code and a web application that classifies the meditative state. But, perhaps most importantly, I provided some recommendations so that machine learning approaches may be more successful in the future. For example, I suggested an approach to acquiring labeled data. This would involve recruiting a sample of meditators, having them complete a survey before meditating (to get demographics, info on their current mental state, etc.), have them meditate with the Zendo Tools app, then have them complete another survey where they describe their meditation experience and when they may have reached peak meditation. Having this labeled data and some additional time-fixed covariates could allow for more impactful analyses with supervised machine learning methods.