Overview
This tutorial will demonstrate how to adjust for selection bias. First, generate a dataset of 100,000 rows with the following binary variables:
- X = Exposure (1 = exposed, 0 = not exposed)
- Y = Outcome (1 = outcome, 0 = no outcome)
- C = Known Confounder (1 = present, 0 = absent)
- S = Selection (1 = selected into the study, 0 = not selected into the study)
set.seed(1234)
n <- 100000
c <- rbinom(n, 1, .5)
x <- rbinom(n, 1, plogis(-.5 + .5 * c))
y <- rbinom(n, 1, plogis(-.5 + log(2) * x + .5 * c))
s <- rbinom(n, 1, plogis(-.5 + 1.5 * x + 1.5 * y))
df <- data.frame(X = x, Y = y, C = c, S = s)
rm(c, x, y, s)
This data reflects the following causal relationship:
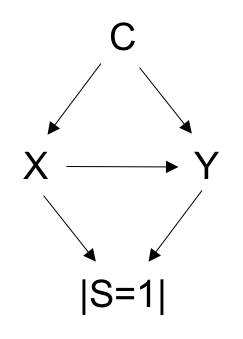
From this dataset note that P(Y=1|X=1, C=c, U=u) / P(Y=1|X=0, C=c, U=u) should equal expit(log(2)). Therefore, odds(Y=1|X=1, C=c, U=u) / odds(Y=1|X=0, C=c, U=u) = ORYX = exp(log(2)) = 2
Compare the biased model to the bias-free model.
nobias_model <- glm(Y ~ X + C,
family = binomial(link = "logit"),
data = df)
exp(coef(nobias_model)[2])
c(exp(coef(nobias_model)[2] + summary(nobias_model)$coef[2, 2] * qnorm(.025)),
exp(coef(nobias_model)[2] + summary(nobias_model)$coef[2, 2] * qnorm(.975)))
ORYX = 2.04 (1.99, 2.09)
This is the value we would expect based off of the derivation of Y.
To represent selection bias, the data for analysis will include only those observations with S=1. We will sample with replacement among those with S=1 from the original dataset so that the sample size remains 100,000.
biased_model <- glm(Y ~ X + C,
family = binomial(link = "logit"),
data = df[sample(seq_len(n), n, replace = TRUE, df$S), ])
exp(coef(biased_model)[2])
c(exp(coef(biased_model)[2] + summary(biased_model)$coef[2, 2] * qnorm(.025)),
exp(coef(biased_model)[2] + summary(biased_model)$coef[2, 2] * qnorm(.975)))
ORYX = 1.32 (1.29, 1.36)
The odds ratio of the effect of X on Y decreases from ~2 to ~1.3 when the analysis is conducted exclusively on participants with S=1. In a perfect world, an investigator would analyze data consisting of the exact same population as the population in which inferences are desired (i.e. the target pop. = source pop.). However, it is often not possible to collect data from the entire source population, and sometimes the selected sample will have collider stratification bias due to the conditional study selection. This bias analysis will allow for the adjustment of selection bias even when data for S is unavailable. We will instead rely on assumptions of how the exposure and outcome affect selection. These assumptions are quantified in bias parameters used in the analysis. Using these bias parameters, we can model the probability of S, which will then be incorporated as a weight in the outcome regression to provide the bias adjustment.
Normally, the values of the bias parameters are obtained externally from the literature or from an internal validation sub-study. However, for this proof of concept, we can obtain the exact values of the parameters. We will perform the final regression of Y on X as if we are only analyzing those with S=1. Since we have the values of S in our data, we can model how S is affected by X and Y to obtain accurate bias parameters.
s_model <- glm(S ~ X + Y, data = df, family = binomial(link = "logit"))
summary(s_model)
These parameters can be interpreted as follows:
- Intercept = log[odds(S=1|X=0, Y=0)]
- X coefficient = log[odds(S=1|X=1, Y=0)] / log[odds(S=1|X=0, Y=0)] i.e. the log odds ratio denoting the amount by which the log odds of S=1 changes for every 1 unit increase in X among the Y=0 subgroup.
- Y coefficient = log[odds(S=1|X=0, Y=1)] / log[odds(S=1|X=0, Y=0)]
Now that values for the bias parameters have been obtained, we’ll use these values to perform the bias adjustment. We’ll build the analysis within a function for quick reiteration. Bootstrapping will be used in order to obtain a confidence interval for the ORYX estimate.
Bias Adjustment
The steps to adjust for selection bias are as follows:
- Sample with replacement from the dataset among rows with with S=1 to get a bootstrap sample.
- Predict the probability of S by combining the bias parameters with the data for X and Y via the inverse logit transformation.
- Using the sampled dataset (bdf), model the weighted logistic outcome regression [P(Y=1)| X, C]. The weight used in this regression is the inverse probability of selection, obtained in the previous step.
- Save the exponentiated X coefficient, corresponding to the odds ratio effect estimate of X on Y.
- Repeat the above steps with a new bootstrap sample.
- With the resulting vector of odds ratio estimates, obtain the final estimate and confidence interval from the median and 2.5, 97.5 quantiles, respectively.
adjust_sel_loop <- function(
coef_0, coef_x, coef_y, nreps, plot = FALSE
) {
est <- vector()
for (i in 1:nreps){
bdf <- df[sample(seq_len(n), n, replace = TRUE, df$S), ]
prob_s <- plogis(coef_0 + coef_x * bdf$X + coef_y * bdf$Y)
final_model <- glm(Y ~ X + C,
family = binomial(link = "logit"),
weights = (1 / prob_s),
data = bdf)
est[i] <- exp(coef(final_model)[2])
}
out <- list(
estimate = round(median(est), 2),
ci = round(quantile(est, c(.025, .975)), 2)
)
if (plot) {
out$hist <- hist(exp(est))
}
return(out)
}
We can run the analysis using different values of the bias parameters. When we use the known, correct values for the bias parameters that we obtained earlier we obtain ORYX = 2.05 (2.01, 2.07), representing the bias-free effect estimate we expect based on the derivation of the data.
set.seed(1234)
correct_results <- adjust_sel_loop(
coef_0 = coef(s_model)[1],
coef_x = coef(s_model)[2],
coef_y = coef(s_model)[3],
nreps = 10
)
correct_results$estimate
correct_results$ci
The output can also include a histogram showing the distribution of the ORYX estimates from each bootstrap sample. We can analyze this plot to see how well the odds ratios converge.

If instead we use bias parameters that are each double the correct value, we obtain ORYX = 3.92 (3.84, 3.96), an incorrect estimate of effect.
set.seed(1234)
incorrect_results <- adjust_sel_loop(
coef_0 = coef(s_model)[1] * 2,
coef_x = coef(s_model)[2] * 2,
coef_y = coef(s_model)[3] * 2,
nreps = 10
)
You can find the full code for this analysis on my github.